BrainStars: 1419083_at: Tnfsf11
| Affymetrix Probe set ID | 1419083_at |
|---|---|
| Gene Name | tumor necrosis factor (ligand) superfamily, member 11 |
| Gene Symbol | Tnfsf11 Ly109l ODF OPG OPGL RANKL Trance |
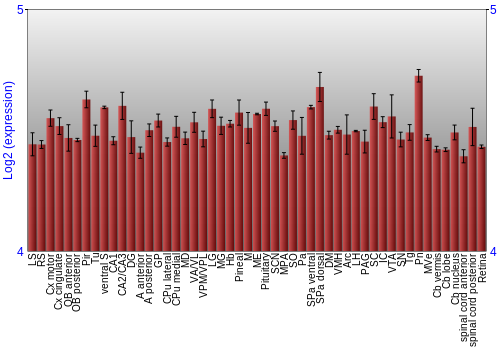
BrainStars Expression at 51 brain regions
PDF files:
Expression Graph,
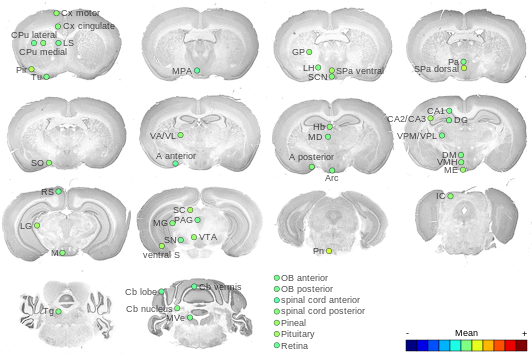
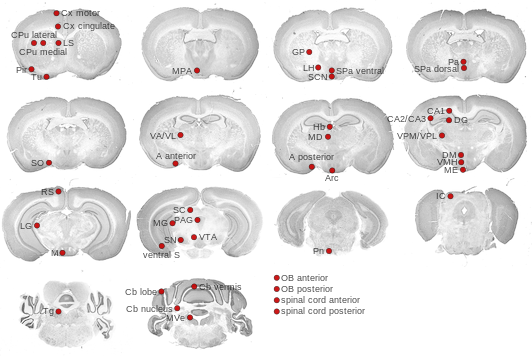
Expression Map
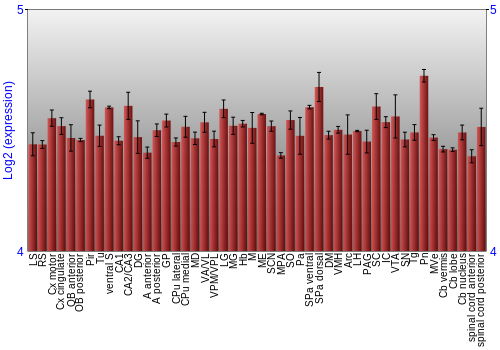
BrainStars classification of 48 brain regions by multi-state gene analysis
| # States | 1 |
|---|---|
| Score | 46.1840016882579 |
PDF files:
Expression Graph,
Expression Map
Other brain expression resources
Cross section images
The ViBrism cross section images (whole brain mode: beta version) were provided by the ViBrism team. A full 3-D model of the expression is available at their ViBrism site.
The ABA expression energy cross section images were obtained from the Allen Mouse Brain Atlas, Allen Instiute for Brain Science © 2008. Please refer to their original terms of use.
Links to other resources
BrainStars 1-D expression files
You can download BrainStars expression data in a table.| TSV format file | BrainStars expression in a tab-separated values format |
| CSV format file | BrainStars expression in a camma-separated values format |
BrainStars 3-D expression files
You can download this entry's BrainStars expression data putting on 3-D mouse brain models.| VCAT format file | BrainStars expression putting on a brain model. The file can be viewed with a software, VCAT, downloadable from VCAD@RIKEN site. |
| XPR format file | BrainStars expression putting on a Allen Reference Atlas brain model. You can view the XPR file with the Brain Explorer software provided by the Allen Institute. |
External links
| Entrez GeneID | 21943 |
|---|---|
| MGI ID | MGI:1100089 |
| Ensembl | ENSMUSG00000022015 |
| UniGene ID | Mm.249221 |
| RefSeq Transcript | NM_011613 |
| RefSeq Protein | NP_035743 |
| Agilent ID | A_51_P465640 A_52_P488947 |
| Allen Brain Atlas | Tnfsf11 |
| ViBrism | Tnfsf11 |
| EMAGE | Tnfsf11 |